iGEM (the international Genetically Engineered Machines Competition) this year consisted of ~130 fantastic teams presenting their summer research over the 3 day competition at MIT. My question for you is, if you’re not at the Jamboree hearing the suggestions of “Check out Harvard’s iGarden”, or “Check out Cambridge reading in the dark by a jug of luminescent bacteria”, how are you supposed to find all the best stuff? My answer is – read this blog.
A bit about me: I was an undergraduate on the Brown iGEM 2007 team, we worked on making a bacteria that would detect lead in a water sample and then glow. After graduating in 2008, I was a judge for the Foundational Advance track in 2009, and the Software track in 2010.
And a necessary disclaimer — iGEM is a magical organization run by a sliver of extremely dedicated staff, and many many volunteers. I am one of those volunteers. The opinions expressed here are mine and do not represent the opinions of iGEM Headquarters in any way.
In 2008 I missed iGEM and tried to filter through some of the 100+ wikis. That takes a long time to read through and you can’t ask the teams questions! Because of this, I’m putting together this set of blog posts over the next week to share the events of iGEM 2010.
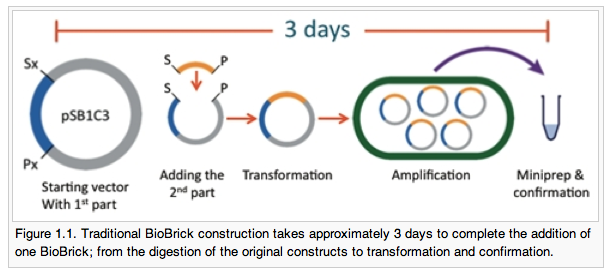
First, let’s talk about Alberta’s BioBytes project. Building DNA is too hard. Are you learning to reprogram E. Coli by putting new DNA into them? In 2007 during my first experience with iGEM, cloning took us 1 month to learn — granted, we were starting from almost zilch wetlab experience and were still getting our lab set up. How about sticking pieces of DNA together – ligation? We spent the last month of our summer on that and never got it to work. I’d be content if the answer was that we were an outlier except as a judge for iGEM in 2009 and 2010 I see several teams falling into the same ruts, and that’s because biology is still hard. Really hard.
Alberta’s project is elegant. Let’s make it easier to build DNA and put it in a cell. What might take a newbie 3 months of daily lab work to tackle, Alberta brought in 5 high school students and they did it in an afternoon – 45 minutes if I recall.
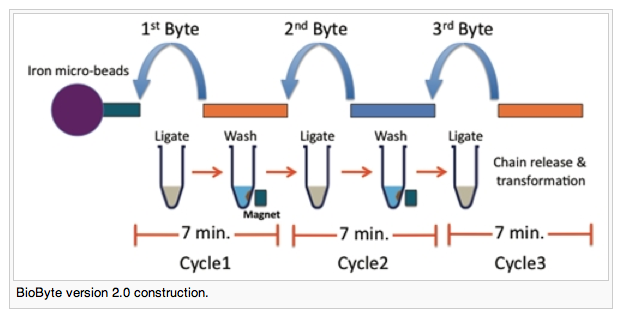
Jumping off of their work done in 2009 to make a rapid way to assemble DNA (earning them the Foundational Advance award), Alberta built a system which allows for the rapid assembly of parts using simple components. It doesn’t even need a “real” pipette to move liquids around! The first byte anchors a small piece of DNA to a magnetic microbead. This serves as the structure upon which each successive “byte” is added — see the diagram below. Wait a few minutes (that’s all you need for ligation), wash with buffer to removes any excess DNA, and then add your next part.
It works and their documentation is clean and simple (
lots of gel electrophoresis). I’ve given you a good summary here, now go read the rest!
Learn more: Alberta put together a sweet “product” website
And Wiki, start with their methods page: